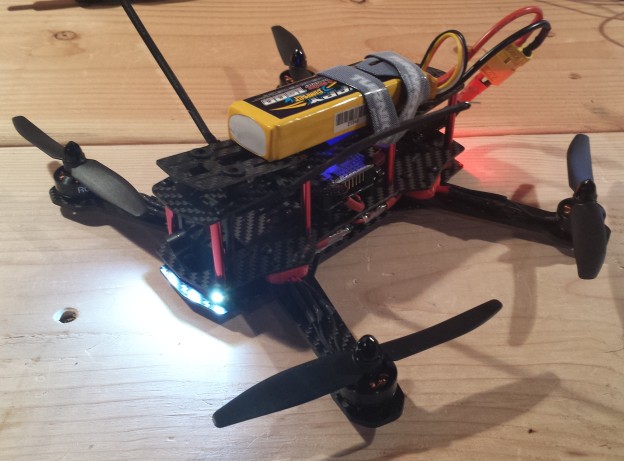
The ZMR250 is a quadcopter frame in the 250mm size class. Compared to the popular DJI Phantom, the platform is significantly smaller, lighter and intended as a racing quad. After starting out with my RCExplorer Tricopter V2 build a few years ago, I kept my eye out for a quadcopter that could easily be packed inside a briefcase for portability. Being small and fast, the ZMR250 excels as an FPV platform for low, fast flying in an affordable and surprisingly durable platform. In the following post, I detail a few key lessons I learned and an overview of my hardware choices.